 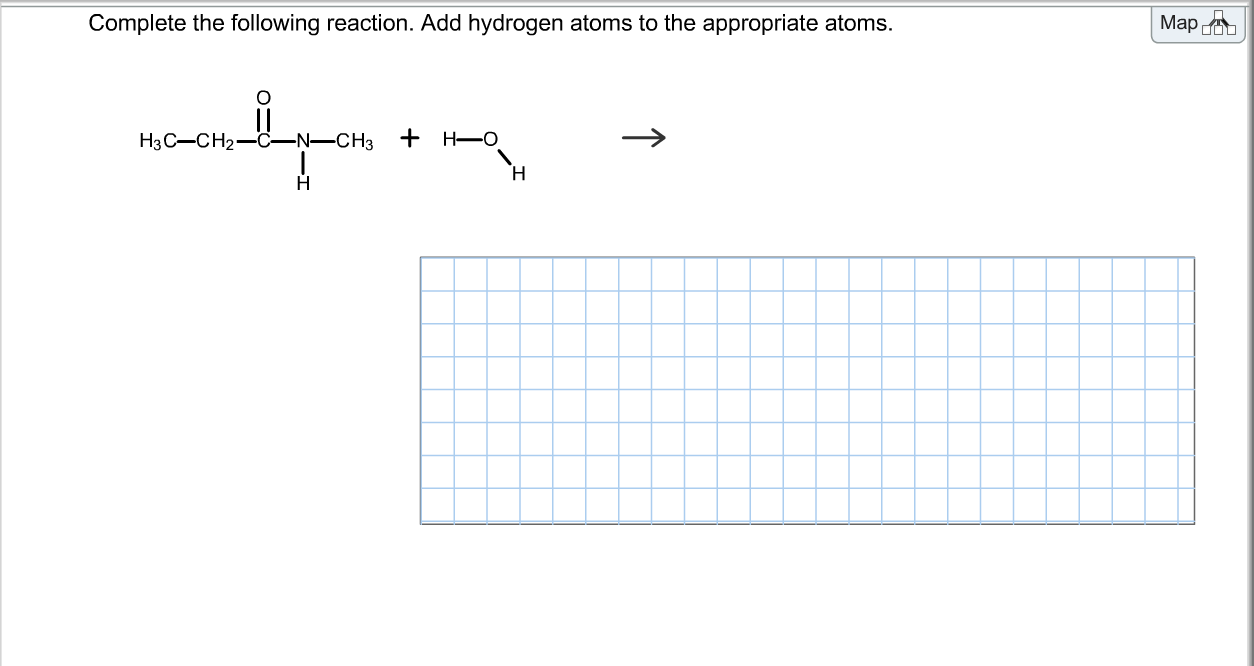
We border a very good time scrutinizing the original output in ChemDoodle. Bonds to heteroatoms or carbons, while implicit hydrogens do not appear directly in the structure or are simply written together with the other atom.Available as ChemDoodle 2D and ChemDoodle 3D. 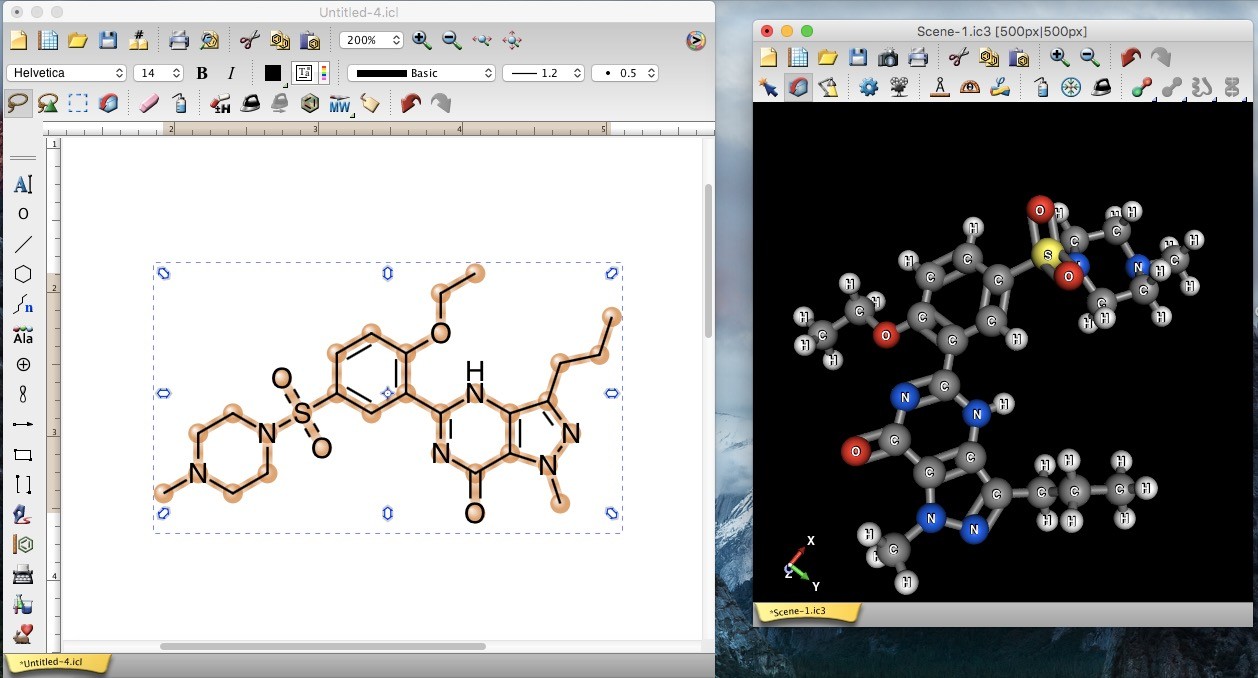
ChemDoodle Web Components (abbreviated CWC, iChemLabs, LLC) is a light-weight (340 KB) JavaScript/HTML5 toolkit for chemical graphics, structure editing, interfaces, and informatics based on the proprietary ChemDoodle desktop software.Provide features like drawing, IUPAC naming, cheminformatics etc. The library uses and WebGL technologies and other HTML5 features to provide solutions for creating chemistry-related applications for the web on. Is your feature request related to a problem? Please describe.Looking to sewer 3D graphics for organizations and chemistry. ChemDoodle 3D v2 is a massive update from the first version included in ChemDoodle 7 and can be used independently from ChemDoodle desktop. Many semiempirical tools warn if explicit hydrogen atoms are missing and add them automatically if needed (Ampac, Spartan, Gaussian etc). Stopping chemdoodle from auto adding hydrogens update# First, load target protein structure (here is 1UBQ), then go to the tool bar, find Tools, select Structure Editing and the popped sub-panel has AddH. Currently xtb calculates any molecule with and without explicit hydrogens added. After clicking AddH, the Add hydrogens window shows. If you have no particular goals, simply click OK button and UCSF Chimera will generate hydrogens for you. This is true for coord (tmol) and mol/sdf input. While we can expect that the user takes care of it, it would be a friendly and helpful feature nevertheless. Plus new users who are not aware of the fact, may just end up with false results. There's also the issue of solvent & counter-ions present in systems like aspartyl proteases with adjacent carboxylates. (R3) Input check, add information if explicit hydrogens are required in the mol/sdf file (xtb does not care about molecules to be chemically valid, they currently can have hydrogens attached or not, xtb will perform an optimization either way) give potential warning if hydrogens are missing. Accessing Through GUI A->hydrogens->add Syntax normal usage hadd (selection) API usage cmd. Same as for 1., we need hydrogen atoms for the calculation, unless we preprocess the structure explicitly such input is not suitable for GFN-xTB calculations. Optimized geometry written to: ordĮxample 2 (xtb benzene with explicit hydrogen): Still gives Energy and HOMO-LUMO, even with hydrogens missing. Gives Energy and HOMO-LUMO, with explicit hydrogens added. Xtb benzene-withhydrogen.tmol -opt extremeĪ simple warning that the solution is not correct in case of explicit hydrogens missing, plus a command line switch to automatically add missing hydrogens. If possible state how you can assist in providing data or code to to implement the feature Use obabel and molconvert to add explicit hydrogens. Stopping chemdoodle from auto adding hydrogens code# I can test molecules, including charged and radical entities.openbabel add explicit hydrogens (from help file). 
Stopping chemdoodle from auto adding hydrogens code#.Stopping chemdoodle from auto adding hydrogens update#. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
